Table of Contents
Therapeutic Targeting of SARS-CoV-2
Created by: Michael D'Ercole, Onela Esho, Jasmine Leung, Meet Patel, and Amandeep Nagi
Introduction
Coronaviruses and SARS-CoV-2
SARS-CoV-2 is the zoonotic coronavirus that causes the disease COVID-19. More specifically, SARS-CoV-2 is a bat coronavirus as it originated in bats and was subsequently passed to humans (Fan et al., 2019). Although the news of a bat-originating virus shocked the general public, the fact that bats are viral reservoirs has been well known in the scientific community (Tang et al., 2020). Coronaviruses are a group of RNA-based viruses that share a common shape, construction, and genome. Coronaviruses are also highly pathogenic to humans and animals and can cause a range of disease states, from upper respiratory infections to enteritis (Fehr & Pearlman, 2015). Bat coronaviruses in particular have been responsible for several recent disease outbreaks affecting both humans and livestock. Including SARS-CoV-2, bat coronaviruses have now caused four major viral outbreaks in the last two decades (Fan et al., 2019).
| Date of initial outbreak | Coronavirus strain | Disease |
| 2003 | SARS-CoV-1 | Severe acute respiratory syndrome |
| 2012 | MERS-CoV | Middle East respiratory syndrome |
| 2016 | SADS-CoV | Swine acute diarrhea syndrome |
| 2019 | SARS-CoV-2 | Coronavirus disease 2019 |
Coronavirus genome
Coronaviruses share a similar positive-sense, single-stranded RNA genome which ranges in size from 27-32 kb (Fan et al., 2019). In the world of RNA viruses, a genome of this size is considered large. In fact, the average genome length of all RNA virus families is only 9 kb (Belshaw et al., 2007). Positive-sense, single-stranded RNA acts as messenger RNA (mRNA) and can be directly translated into protein by host cell ribosomes. The viral genome also acts as a template for RNA replication and encodes its own RNA-dependent RNA polymerase (RdRP) (Ahlquist et al., 2003). The genomes of all coronaviruses are common in their organization and allocation of genetic code (Fan et al., 2019). Two-thirds of the genome is generally allocated to the ORF1ab gene which codes for two overlapping polyproteins, ORF1a and ORF1b (Fan et al., 2019). ORF1a and ORF1b are cleaved to produce an array of non-structural proteins. Non-structural proteins are those involved in the replication of viral components, processing of new viral particles, and the suppression of host immune responses. Examples of non-structural proteins include RdRP, ribose-methyltransferase, and helicase (NCBI, 2021). The remaining third of the genome usually encodes for structural and accessory proteins. Structural proteins constitute the coronavirus virion and are primarily involved in receptor binding and viral fusion. Accessory proteins vary the most between viral subgroups and are the main determinant of genome length (Fan et al., 2019).
Coronavirus structure
Coronavirus virions have a characteristic spherical shape with spike-like projections that protrude from the viral envelope. Under a microscope these projections resemble a stellar corona, hence the name coronavirus (Fehr & Pearlman, 2015). Coronavirus virions are comprised of four structural proteins:
Spike (S): Homotrimers of the S protein form club-shaped spike projections that protrude from the viral envelope. These spike projections mediate attachment of the virus to host cells via an interaction with host cell receptors (Fehr & Pearlman, 2015).
Envelope (E): The E protein of coronaviruses are often dissimilar and scarce but share an overall common architecture. Consequently, they have not been accurately modelled and are not well understood. Current data suggests that the E protein is a transmembrane protein involved in virion assembly and release (Fehr & Pearlman, 2015).
Membrane (M): M protein dimers are found in high quantities within coronavirus virions and organize themselves to form the characteristic spherical shape of coronaviruses (Fehr & Pearlman, 2015).
Nucleocapsid (N): The N protein contains two RNA-binding domains, the N-terminal, and C-terminal domain. Both domains have been shown to bind viral RNA and assist in the packaging of the viral genome into new viral particles. The viral nucleocapsid consists entirely of N proteins (Fehr & Pearlman, 2015).
SARS-CoV-2 and the role of the spike protein
To gain entry into human cells, SARS-CoV-2 targets angiotensin-converting enzyme 2 (ACE2), a transmembrane receptor found abundantly in the epithelia of the lungs, small intestine, kidneys, heart, arteries, and veins (Shereen et al., 2020; Hamming et al., 2004). The binding of the virus to host cells is mediated by the S protein homotrimers that protrude from the viral envelope. The S protein of SARS-CoV-2 can be divided into three units:
- Signal peptide [residues 1-13].
- S1 subunit [residues 14-685].
- S2 subunit [residues 686-1273].
The S1 subunit contains a receptor-binding domain (RBD) which has been shown to take on an opened and closed conformation. In its open conformation, the RBD binds ACE2 and initiates viral fusion by triggering the formation of endosomes. The fusion of the viral and host cell membranes is supported by the S2 subunit. The S2 subunit helps by anchoring the virus to the host cell membrane and disrupting the host cell’s lipid bilayer. As the first point of contact between the virus and its host cells, the S protein is a prime target for prophylactic and post-exposure therapies (Huang et al., 2020).
Vaccines and the need for alternative viral therapies
To control the spread of COVID-19, the Canadian government has imposed rules and regulations which seek to minimize in-person interactions. However, social distancing, the use of masks, and the closure of schools and businesses are short-term solutions to a long-term problem. Vaccines are often the most talked-about method of combating viral outbreaks, and for good reason. Vaccines offer a long-term solution by training our cells to fight off a specific pathogen (CDC, 2020). But, as with any form of medical treatment or preventative, vaccines do not come without their limitations. For one, vaccines only offer prophylactic protection, and thus the administration of a vaccine will not help an infected individual (CDC, 2018). Additionally, the protection provided by a vaccine does not take effect immediately, and in the case of COVID-19 mRNA vaccines, the administration of more than one dose may be required (CDC, 2018; Public Health Ontario, 2020). Thus, there is a need for alternative therapeutic options that provide rapid protection against viral infection or alleviate disease symptoms.
Small molecules and immunotherapeutics may be the solution to this problem. The repurposing of approved drugs, such as steroids and antivirals, is a rapid approach to post-exposure therapy (Wu et al., 2021). Since the pharmacodynamic and pharmacokinetic properties of approved drugs is already known, the time and money required to repurpose a drug is much less than that required to develop a new drug (Gupta, 2020). For example, the commonly used corticosteroid dexamethasone has been shown to reduce the mortality of COVID-19 patients on ventilators by up to 33% (WHO, 2020). Additionally, monoclonal antibodies have gained notoriety as both a from of prophylaxis and post-exposure therapy (Pinto et al., 2020).
Therapeutic Targeting
Antibodies
Monoclonal antibodies
The immune system acts as a defense against foreign infectious agents that cause various forms of disease. The two main parts of the immune systems are the innate immune system and the adaptive immune system. The innate immune system is an immediate, non-specific response against invading pathogens. The adaptive immune system is antigen-specific and comes into effect over time. Two major components of the adaptive immune system are the humoral (antibody-mediated) and cellular (cell-mediated) immune responses (Mahmuda et al., 2017). The humoral immune response comprises of B-lymphocytes which recognize the type of foreign invading antigens and produce specific antibodies against them (Mahmuda et al., 2017). Antibodies are characterized by their specificity to an antigen and ability to consistently neutralize that specific antigen. Considering the unique properties of antibodies, scientists have developed techniques to produce antibodies in vitro, which are referred to as monoclonal antibodies (mAbs).
Antibodies are Y-shaped in structure and consist of four polypeptides: two heavy chains and two light chains. This structure allows the antibody to undergo antigen binding, as well as mediate biological activity. The fragment antigen-binding (Fab) region binds to the antigens. The fragment crystallizable region (Fc) region mediates biological activities such as interacting with cell surface receptors.
Targets of mAbs:
(Manis, 2020)
- Cell Surface Antigen - mAbs bind to cell surface antigens to either block their function or recruit other immune cells and molecules (such as complement) which promote the killing of target cells.
- Plasma Proteins - mAbs bind to plasma proteins to sequester them away from their ligands.
Therapeutic uses of mAbs:
- Cancer therapy - mAb treatment has been approved for several cancers including brain, breast, colon, lymphoma, etc.
- Mechanisms include (Kimiz-Gebologlu et al., 2018):
- Inhibition of factors and receptors that activate the signal pathways used by the cancer cells in division and angiogenesis (e.g., Alemtuzumab).
- Antibody-dependent cellular cytotoxicity (ADCC) (e.g., Rituximab).
- Complement-dependent cytotoxicity (CDC) (e.g., Rituximab).
- Autoimmune diseases - mAb treatment effective against rheumatoid arthritis, Chron’s disease, and ulcerative colitis include Infliximab and Adalimumab which bind to tumor necrosis factor (TNF), TNF-α, and Interleukin-2 (IL-2) on activated T-cells to suppress the immune response (Mahmuda et al., 2017).
- Infectious diseases - Several mAbs are in clinical development for the prevention and treatment of infectious viral diseases such as influenza, human immunodeficiency virus (HIV), ebola, zika, rabies, hepatitis B (HBV), dengue (Salazar et al., 2017), SARS-CoV, and SARS-CoV-2. Recently, the FDA authorized Balanivimab as well as Casirivimab and Indevimab together for emergency use in the treatment of SARS-CoV-2.
Advantages to mAbs:
(Nagarajan et al., 2014; Cheriyedath, 2018)
- mAbs have a high-specificity for a single epitope of an antigen
- mAbs can be renewably generated once their candidate hybridoma or Chinese hamster ovary (CHO) cells have been developed
- mAbs are homogenous and highly consistent
- mAbs can be used for the immediate treatment of a SARS-CoV-2 infection
Disadvantages to mAbs:
(Nagarajan et al., 2014; Cheriyedath, 2018)
- mAbs mono-specificity limits their application
- Minor changes in the antigen’s epitope structure can drastically reduce the binding affinity of mAbs
- The production of mAbs is an expensive process
- The protection from mAbs is short-term (weeks to months)
Neutralizing antibodies
A specific subclass of antibodies, known as neutralizing antibodies (nAb), is essentially responsible for defending cells from pathogens, in its most simplified function. They are produced naturally by the body as part of its humoral response of the adaptive immune system against viruses, microbial toxin, and other pathogens (Cheedarla, 2019). Therefore, their production is triggered by both infections and vaccinations against infections (Salazar et al., 2017). This subclass of antibodies can result in lifelong immunity to certain infections and can even be used to see if a person has developed immunity to an infection after they have recovered from it (Salazar et al., 2017).
Function
Neutralizing antibodies inhibit the infectious capacity of a pathogen by bind in a manner to interact with the surface receptors of their target and disrupting interactions needed for cell entry in to the host cell, or even preventing crucial surface structures from undergoing conformational changes required for entry (Salazar et al., 2017). When considering viral infections, nAbs are capable of binding to glycoprotein tips of enveloped viruses and statically interfering with the pathogen’s ability to recognize and communicate with the host cell’s surface receptors (Walls et al., 2020). And in some cases, the pathogen is rendered incapable of infecting host cells even after the nAbs has dissociated from the virus (Salazar et al., 2017).
Production and transfer
Like other antibodies, nAbs are produced and secreted by B cells. But nAbs can be passively transferred before or after viral infection as either a prophylactic measure, or as a treatment (Jiang et al., 2020). Whereas, vaccines are able to actively stimulate the immune system into creating antigen-specific neutralizing antibodies (Jiang et al., 2020). However, therapeutic nAbs generally only exist in the body for a short range of time, while their treatment efficacy can be influenced by a variety of factors (Jiang et al., 2020). Therefore, neutralizing antibodies with high specificity and strong affinity to their target antigen, along with low toxicity are used in treatments of viral infections (Jiang et al., 2020).
Neutralization of SARS-CoV-2
Passive administration of monoclonal antibodies could have a significant impact on providing an immediate protection measure against SARS-CoV-2, while simultaneously complimenting the development of other prophylactic vaccines (Pinto et al., 2020). As stated, the SARS-CoV-2 spike (S) glycoprotein is responsible for promoting entry into host cells. More specifically, the S protein homotrimers contain a receptor binding domain which binds to the ACE2 receptor on the host cell. Therefore, consequently, the S glycoprotein of SARS-CoV-2 serves as a primary target for therapeutic agents, like nAbs, to be designed around (Pinto et al., 2020).
In a previous study, several human monoclonal antibodies of interest, in terms of neutralizing antibodies that target the host ACE2 receptor-binding domain of SARS-CoV-2, were identified from the memory B cells of a previous SARS infection survivor (Barnes et al., 2020). As a brief summary, memory B cells form following an infectious illness and their lineage is capable of lasting for life (Jiang et al., 2020). They function to remember a pathogen, or an analogue, that has previously infected the host, and launch an antibody defense against a re-infection (Pinto et al., 2020). To characterize the potential cross-reactivity of these antibodies with SARS-CoV-2, a recent study performed a memory B cell screening using peripheral blood mononuclear cells collected from the same patient (Pinto et al., 2020). The study shows further evaluation of the mAbs for binding to the SARS-CoV-2 and SARS-CoV SB domains, as well as to ectodomain trimers of various SARS-CoV variant S glycoproteins (Pinto et al., 2020).
The study found a particular IgG antibody, called S309, was able to successfully bind to both the immobilized SARS-CoV-2 SB domain, as well as to the ectodomain trimer of the S glycoprotein with significant affinities (Pinto et al., 2020). The researchers then performed binding assays to further investigate the mechanism of S309-mediated neutralization. Results indicate that one or more IgG-specific mechanisms might be involved, including (Pinto et al., 2020):
- Cross-linking of S-glycoprotein trimer
- Steric hindrance
- Aggregation of virions
Any combination of these bivalent mechanisms may contribute to the ability of S309 to fully neutralize pseudovirions (Pinto et al., 2020). The study indicates imaging tests were conducted afterwards, specifically cryo-electron microscopy, with results showing that the S309 antigen-binding fragment engages an epitope distinct from the receptor-binding motif, containing a glycan that is conserved within the Sarbecovirus subgenus without competing with ACE2 binding of the antigen (Pinto et al., 2020). Therefore, the study concludes that S309 potentially neutralizes SARS-CoV-2 and SARS-CoV-2 pseudoviruses (Pinto et al., 2020).
Small Molecules
Small molecules are organic compounds characterized by their low molecular weights and have historically been used in the field of medicine and pharmaceuticals (Ngo et al., 2018). Overall, the use of these molecules allows for modulation of various biochemical pathways and provides therapeutic options for various diseases, such as cancer (Ngo et al., 2018). Some of the benefits provided from the use of small molecules as therapeutics are (Ngo et al., 2018):
- Their simple chemical composition makes them easy to manufacture
- Can test multiple variants of molecules through high throughput screening and determine the most effective one
- Typically have stable structures, allowing them to be administered orally
- More affordable compared to alternative therapeutics (e.g. biologics)
However, small molecules are not always the most effective course of disease treatment. Some of the limitations found with their use include (Ngo et al., 2018):
- Cancer cells, bacteria and viral targets may develop mechanisms for drug resistance (e.g. drug inactivation, efflux pumps)
- Limited function in that these molecules typically only target one protein, cell receptor or cellular pathway
Small molecules as potential therapeutics for SARS-CoV-2
The use of small molecules to prevent viral transcription and replication has long been investigated as an attractive option to prevent infection (Hermann, 2016). Prior to designing and synthesizing therapeutic compounds, researchers must have both thorough knowledge of the structural components of the virus in question as well as a deep understanding of essential mechanisms for the virus' survival within a host (Hermann, 2016). With this in mind, early research conducted on SARS-CoV-2 focused largely on understanding the essential proteins and processes that the virus needs to both infect the host cell and produce its viral particles (Fan et al., 2019).
Since there were many studies discussing the structure and pathology of the pre-existing SARS-CoV virus, those findings assisted in the accelerated research done on SARS-CoV-2 in response to the current COVID-19 pandemic (Gordon et al., 2020). In particular, the degree of similarity between the genome of SARS-CoV and SARS-CoV-2 allows researchers to investigate how effective pre-existing small molecules for SARS-CoV would be in preventing SARS-CoV-2 replication (Gordon et al., 2020). Currently, studies are focusing on designing potential inhibitors for many of the different processes that SARS-CoV-2 requires for replication and infection within the host cell (Wu et al., 2020).
Although there are many protein targets to choose from, clinical trials are seeing the most promising results with small molecules that inhibit SARS-CoV-2 RNA-dependant RNA polymerase (RdRp), cysteine main protease (Mpro) and the interaction between SARS-CoV-2 spike protein and ACE-2 host cell receptors (Wu et al., 2020). These proteins serve a different function from one another but are both essential for a successful replication cycle for the SARS-CoV-2 virus (Wu et al., 2020). Some of the small molecules being investigated in current clinical trials for their ability to inhibit these SARS-CoV-2 proteins include (Wu et al., 2020):
1. Nucleoside analogs that inhibit SARS-CoV-2 RdRp
2. Compounds that inhibit SARS-CoV-2 Mpro
3. Small molecules that prevent spike protein (S) and ACE-2 receptor interaction
Even though these drugs have shown promising results, they are still undergoing further testing for their safety and efficacy, or are awaiting FDA approval prior to distribution (Wu et al., 2020). Due to this lengthy process, patients with severe COVID-19 infections continue to suffer with a multitude of complications, one of the most severe being acute respiratory distress syndrome (ARDS) (Prescott et al., 2020). As such, studies are being done to determine the most effective course of treatment to prevent severe disease pathology (Prescott et al., 2020). One of the possible treatments being researched includes (Prescott et al., 2020):
4. Corticosteroids to reduce inflammation
With so many options of small molecule treatments, researchers must prioritize their focus on those that (Wu et al., 2020):
- Are most cost-efficient to make
- Have stable chemical structures for both ingestion and storage purposes
- Have the lowest risk of adverse effects
Nucleoside analogs as RdRp inhibitors
The RdRp encoded by SARS-CoV-2 allows the virus to both replicate its genome and to produce all of the structural components needed for the making of viral progeny (Ahlquist et al., 2003). As such, it serves an essential role for the virus' ability to spread from one host cell to another and cause disease (Ahlquist et al., 2003). By inhibiting the polymerase activity of this protein, you are essentially preventing viral spread and are reducing the chance of severe COVID-19 disease (Wu et al., 2020).
How it works:
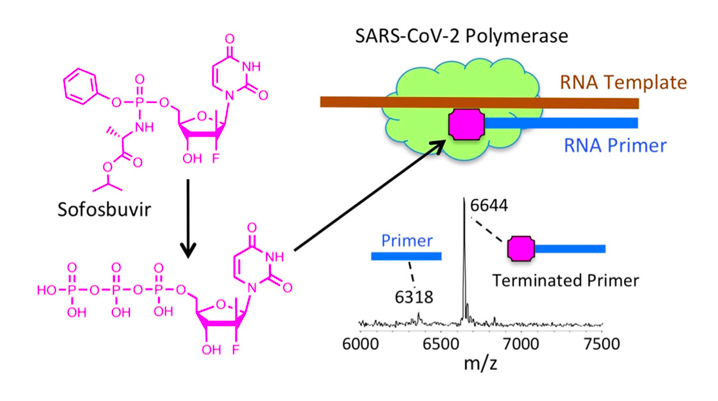
Nucleoside analogs have been known to inhibit the RdRp of SARS-CoV and MERS-CoV (Chien et al., 2020). After discovering that the RdRp of SARS-CoV and SARS-CoV-2 have a 98% similarity in their nsp12 region (site of polymerase activity), the researchers hypothesized that similar drugs can be used for COVID-19 as those used for SARS (Chien et al., 2020). These nucleoside analogs work by first being introduced into the body as a prodrug and are transformed into an active nucleoside triphosphate form once inside of a virally-infected cell (Chien et al., 2020). For example, prodrugs like Remdesivir and Sofosbuvir are activated and able to successfully inhibit the RdRp of SARS-CoV, MERS-CoV and SARS-CoV-2 (Chien et al., 2020). In its active form, the compound is able to bind to the nsp12 region of RdRp and inhibit its polymerase activity (Chien et al., 2020). This prevents SARS-CoV-2 from replicating its genome and synthesizing necessary components for viral spread during a COVID-19 infection (Chien et al., 2020).
Limitations:
- Conversion from prodrug to active form can produce some toxic metabolites depending on the enzymatic reaction (Hajdo et al., 2010).
- May not be delivered to the specific target cells (e.g. lung cells infected with SARS-CoV-2) (Hajdo et al., 2010).
Mpro (main protease) inhibitors
Viruses of the coronavirus family (SARS-CoV, MERS-CoV and SARS-COV-2) have a highly conserved structure of Mpro, a cysteine protease which functions to modify polypeptides produced post-translationally (Jin et al., 2020). One of its main protein modifications is that of the pp1a and pp1ab region of the replicase enzyme (Jin et al., 2020). It is apparent through research that Mpro is essential to the virus' life cycle and that its structure differs from that of human Mpro (Jin et al., 2020). As such, Mpro serves as an attractive target for small molecule inhibitors (Jin et al., 2020).
How it works:
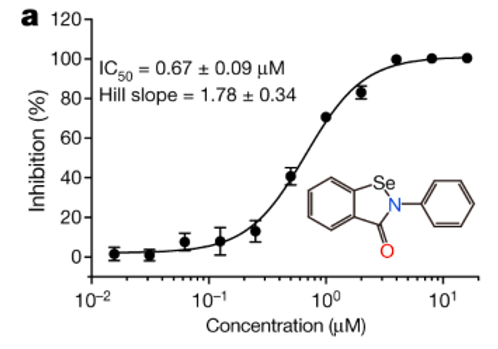
High throughput screening methods tested for 10,000 different drug candidates to determine which one is the most efficient inhibitor of Mpro (Jin et al., 2020). It was determined that ebselen, an organoselenium compound, was most efficient for covalent bonding to inhibit Mpro and showed the strongest antiviral effects overall (Jin et al., 2020). With these findings, the researchers show that repurposing of drugs used for SARS-CoV and MERS-CoV Mpro allows for successful inhibition of SARS-CoV-2 Mpro (Jin et al., 2020). Furthermore, the difference in SARS-CoV-2 and human Mpro homology ensures that the chances of adverse effects of ebselen are limited (Jin et al., 2020).
Limitations:
- The active sites of Mpro for SARS-CoV and SARS-CoV-2 are different in structure, so cannot repurpose drugs used for SARS (Bzówka et al., 2020).
- Mpro is susceptible to mutations, making small molecule design challenging (Bzówka et al., 2020).
Blocking spike (S) protein- ACE-2 receptor interaction
SARS-CoV-2 uses its spike (S) protein to bind to host cell ACE-2 receptors and facilitate entry into the cell (Shereen et al., 2020). As such, blocking this essential protein interaction is one of the most researched methods of preventing SARS-CoV-2 infection and many therapeutic candidates are being considered (Shereen et al., 2020). Although neutralizing antibodies are the primary candidate for this purpose, small molecules are also being designed with the same purpose (Yi et al., 2004)
How it works:
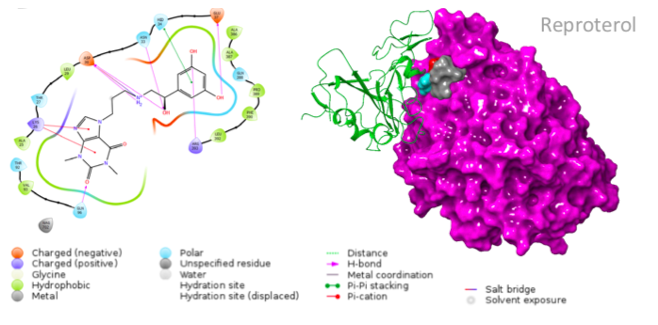
The small molecules candidates designed for this reaction are different in structure from one another, and thus bind to different residues of the ACE-2 receptor (Smieszek et al., 2020). A drug that shows promising results is Reproterol, a β2 adrenoreceptor agonist that has been successfully used for treatment of asthma symptoms (Smieszek et al., 2020). It works by binding to the His34 or Asp30 residues of the ACE-2 receptor and is therefore able to inhibit the interaction between this receptor and the S protein of SARS-CoV-2 (Smieszek et al., 2020).
Limitations:
- More research is being done on other potential therapeutics (e.g. the use of monoclonal antibodies) (Pinto et al., 2020).
- Has not been tested extensively (Smieszek et al., 2020).
Corticosteroids to reduce inflammation in severe COVID-19 cases
Studies have shown that COVID-19 patients with the highest mortality rates are those that experience a cytokine storm (Ragab et al., 2020). A cytokine storm is characterized by excessive production of pro inflammatory cytokines, such as IL-1, IL-6 and TNF- α (Ragab et al., 2020). Although these cytokines are necessary for induced killing of virally-infected cells, their overproduction can cause tissue damage, leading to multi-organ failure and subsequent death (Ragab et al., 2020). For such severe cases, health care workers may benefit through administering anti-inflammatory drugs, such as corticosteroids (Prescott et al., 2020).
How it works:
Corticosteroids are compounds with anti-inflammatory function, and are prescribed for many conditions such as asthma, chronic obstructive pulmonary disease (COPD) and irritable bowel disease (IBD) (Prescott et al., 2020). The use of corticosteroids like hydrocortisone and dexamethasone have shown the ability to decrease symptoms of patients with acute respiratory distress syndrome (ARDS) (Prescott et al., 2020). Since development of ARDS is common in severe COVID-19 disease, these corticosteroids may serve as an effective means of preventing multi-organ failure (Prescott et al., 2020).
Limitations:
- Findings from studies done on corticosteroid use for COVID-19 treatment have been inconsistent (Prescott et al., 2020).
- Secondary infections may occur following corticosteroid treatment, since they reduce immune system function overall (Prescott et al., 2020).
Other Immunotherapies
Immunotherapy is the field of immunology that aims to identify treatments for diseases through induction, enhancement, or suppression of an immune response, in order to achieve the therapeutic effect (Wraith, 2017).
Two main areas for immunotherapies
(Wraith, 2017)
- Activating immunotherapies for cancer by enhancing the patients' immune response
- Suppressive immunotherapies for autoimmune disease by repressing the immune response
Common types of immunotherapy
(Immunotherapy - StatPearls - NCBI Bookshelf, n.d.)
- Drugs (e.g., immunosuppressors)
- Biologicals (e.g., cytokines, monoclonal antibodies, and antisera)
- Transplantation (e.g., bone marrow)
- Immunizations (e.g., prophylactic and therapeutic vaccines)
Other types of immunotherapies
(Pashaei & Rezaei, 2020)
- Anti-inflammatory therapies
- Passive immunotherapy
- Convalescent plasma treatment
- Intravenous immunoglobulin (IVIG therapy)
- Mesenchymal stem cells (MSCs) therapy
Limitation of immunotherapy
(Immunotherapy - StatPearls - NCBI Bookshelf, n.d.)
- Side effects
- Partial response (does not work for everyone)
- May harm organs and system (drugs cause the immune system to attack organs)
- May stop having an effect over time
Anti-inflammatory therapy
Anti-inflammatory drugs limit inflammatory processes that drive the worsening of the disease (Pashaei & Rezaei, 2020).
Example for SARS-Cov-2
(Pashaei & Rezaei, 2020)
- Important cytokine: IL-1β released during pyroptosis and acts through autocrine stimulation of tissue macrophages leading to further inflammatory cytokines production (Pashaei & Rezaei, 2020)
- Recombinant IL-1 receptor antagonist (rIL-1Ra, Anakinra)
- Immunosuppressive drug
- Blocks the binding of both IL-1α and IL-1β to the IL-1 receptor
- Inhibits IL-1 pro-inflammatory effects.
Limitation
(King et al., 2020)
- Increased rates of infection were reported with prolonged anakinra use in combination with TNF-α blockade, but not with short-term use
- Only target treatment to individuals considered to have hyperinflammation
- Carries the risk of harm in the wrong patient group by potentially targeting beneficial inflammation
Passive therapy
I. Convalescent plasma treatment
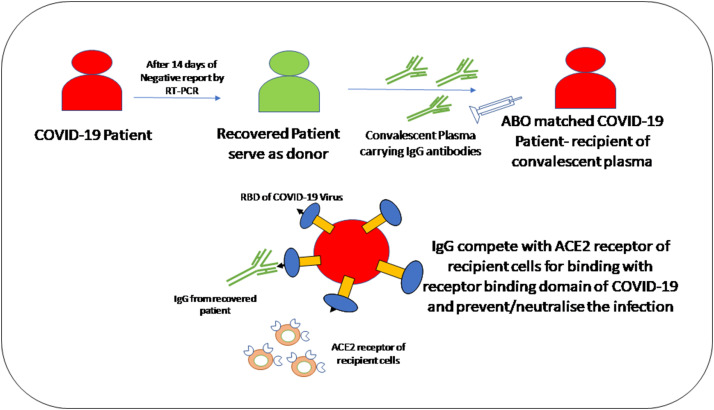
Convalescent plasma treatment is a passive polyclonal antibody administration to supply immunity against viral infections and improves survival rate, which is called passive immunotherapy (Pashaei & Rezaei, 2020).
Example for SARS-CoV-2
(Pashaei & Rezaei, 2020)
- Convalescent plasma treatment provides an immunomodulatory effect via inhibition of macrophage activation and cytokine storm.
- Studies on COVID-19 patients have shown the presence of antibodies against SARS-CoV-2 in serum such as IgG, IgM, and IgA with varying sensitivities.
- Duan et al. had shown infusion of 200 ml of convalescent plasma containing viral neutralizing antibodies with titers more than 1:640 to 10 patients with COVID-19 improved their clinical symptoms and decreased CRP level in all patients.
Limitation
(Nagoba et al., 2020)
- Immunological reactions e.g. severe allergic reaction
- Risk of reinfection
II. Intravenous immunoglobulin (IVIG therapy)
IVIG is a product made from a fractionation of pools of thousands of plasma donations collected in blood transfusion services. Traces of IgM and IgA and cytokines are present in IVIG. IVIGs have several proposed mechanisms of action to achieve their therapeutic effects (Pashaei & Rezaei, 2020).
Example for SARS-CoV-2
(Pashaei & Rezaei, 2020)
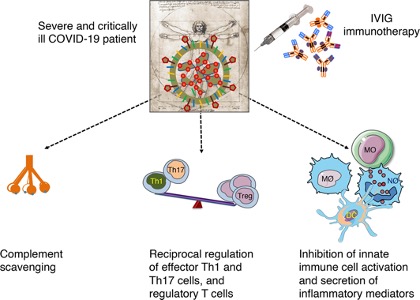
- IVIG therapy induced passive immunity with anti-inflammatory and immunomodulatory effects.
- IVIG blocks the activation and release of inflammatory cytokines from innate immune cells, inhibits Th1 and Th17, which are involved in inflammation, probably enhances regulatory T (Treg) cells as an immune suppressor cell for decreasing inflammation, which is indicated a significant reduction in the number of peripheral blood Treg cells in severe cases of patients with COVID-19.
- Study shows patients decreased hospital stay and ventilator use and improved clinical symptoms.
Figure 13: IVIG mechanism of action in COVID-19: IVIG targets inflammatory process and cytokine storm by complement scavenging; reciprocal regulation of effector Th1 and Th17 cells, and regulatory T cells (Treg); and inhibition of innate immune cell activation and secretion of inflammatory mediators (Galeotti et al., 2020).
Limitation
(Guo et al., 2018)
- Immediate adverse effect
- Flu-like symptoms
- Delayed adverse effect
- Risk of having neurological disorder
- Can be lethal
- Only affect less than 1% of patients
Mesenchymal stem cells (MSCs) therapy
Mesenchymal stem cells (MSCs) are multipotential stem cells that are recognized via self-renewal capacity, generation of clonal populations, and multilineage differentiation. MSCs are present in nearly all tissues of the body, playing an essential role in the repair and generation of tissues (Kavianpour et al., 2020).
MSCs have broad immunoregulatory properties through the interaction of immune cells in both innate and adaptive immune systems, leading to immunosuppression of many effector activities (Kavianpour et al., 2020).
Example for SARS-Cov-2
(Pashaei & Rezaei, 2020)
- MSCs can reduce the cytokine storm through secretion of immunomodulatory factors or direct interaction with immune cells.
- After transplantation of MSCs, these cells improve the lung function of patients with COVID-19 by triggering endogenous repair and recovering the pulmonary microenvironment due to their regenerative traits.
Limitations
(Durand et al., 2020)
- Potential tumorigenicity
- Currently no pre-clinical efficacy data for the use of MSC in animal models of COVID-19 pneumonia
- Recapitulation of efficacy data obtained in animal models in human subjects remains a major barrier to the successful translation of MSC therapy in clinical settings
Conculsion
Table 1: Summary of comparism between therapeutics which can be used to target SARS-CoV-2
In conclusion, Covid-19 is still an ongoing concern. In the last year, a few treatments have been developed and approved in the form of mRNA vaccines that are approximately 90-95% effective. Until a vaccine for SARS-CoV-2 is developed, we can potentially use therapeutics such as mAbs, small molecules, and others to prevent and treat a SARS-CoV-2 infection. Although, these therapeutics don't offer long-term protection like vaccines, they can still be used to achieve herd immunity. This would essentially lower the spread of the virus by breaking any chains transmission. Therefore, we hope everyone will opt-in to get their vaccines, because you won't be only saving your own lives, but also the lives of others.
Presentation Slides
References
Ahlquist, P., Noueiry, A. O., Lee, W., Kushner, D. B., & Dye, B. T. (2003). Host Factors in Positive-Strand RNA Virus Genome Replication. Journal of Virology, 77(15), 8181-8186. doi:10.1128/jvi.77.15.8181-8186.2003
Alexander, M. R., Sanders, R. W., Moore, J. P., & Klasse, P. J. (2015). Virion aggregation by neutralizing and nonneutralizing antibodies to the HIV-1 envelope glycoprotein. AIDS research and human retroviruses, 31(11), 1160-1165.
Aronson, J. K. (2020, March 25). Coronaviruses – a general introduction [Digital image]. Retrieved January 22, 2021, from https://www.cebm.net/covid-19/coronaviruses-a-general-introduction/
Barnes, C. O., Jette, C. A., Abernathy, M. E., Dam, K. M. A., Esswein, S. R., Gristick, H. B., … & Bjorkman, P. J. (2020). SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature, 1-6.
Belshaw, R., Pybus, O. G., & Rambaut, A. (2007). The evolution of genome compression and genomic novelty in RNA viruses. Genome Research, 17(10), 1496-1504. doi:10.1101/gr.6305707
Bzówka, M., Mitusińska, K., Raczyńska, A., Samol, A., Tuszyński, J. A., & Góra, A. (2020). Structural and evolutionary analysis indicate that the SARS-CoV-2 Mpro is a challenging target for small-molecule inhibitor design. International journal of molecular sciences, 21(9), 3099.
CDC. (2018, July). Understanding How Vaccine Work. Retrieved January 21, 2021, from https://www.cdc.gov/vaccines/hcp/conversations/downloads/vacsafe-understand-color-office.pdf
CDC. (2020, December 28). Understanding mRNA COVID-19 Vaccines. Retrieved January 22, 2021, from https://www.cdc.gov/coronavirus/2019-ncov/vaccines/different-vaccines/mrna.html#:~:text=COVID%2D19%20mRNA%20vaccines%20are,protein%20piece%20on%20its%20surface.
Cheedarla, N., & Hanna, L. E. (2019). Functional and Protective Role of Neutralizing Antibodies (NAbs) Against Viral Infections. In Recent Developments in Applied Microbiology and Biochemistry (pp. 83-93). Academic Press.
Cheriyedath, S. (2018). Monoclonal Antibodies. News Medical Life Sciences. Retrieved from https://www.news-medical.net/life-sciences/Monoclonal-Antibodies.aspx
Chien, M., Anderson, T. K., Jockusch, S., Tao, C., Li, X., Kumar, S., … & Ju, J. (2020). Nucleotide analogues as inhibitors of SARS-CoV-2 polymerase, a key drug target for COVID-19. Journal of Proteome Research, 19(11), 4690-4697.
Durand, N., Mallea, J., & Zubair, A. C. (2020). Insights into the use of mesenchymal stem cells in COVID-19 mediated acute respiratory failure. Npj Regenerative Medicine, 5(1), 1–9. https://doi.org/10.1038/s41536-020-00105-z
Fan, Y., Zhao, K., Shi, Z., & Zhou, P. (2019). Bat Coronaviruses in China. Viruses, 11(3), 210. doi:10.3390/v11030210
Fehr, A. R., & Perlman, S. (2015). Coronaviruses: An Overview of Their Replication and Pathogenesis. Coronaviruses, 1282, 1-23. doi:10.1007/978-1-4939-2438-7_1
Galeotti, C., Kaveri, S. v, & Bayry, J. (2020). Intravenous immunoglobulin immunotherapy for coronavirus disease‐19 (COVID‐19). Clinical & Translational Immunology, 9(10), e1198. https://doi.org/10.1002/cti2.1198
Gordon, D. E., Jang, G. M., Bouhaddou, M., Xu, J., Obernier, K., White, K. M., . . . Krogan, N. J. (2020, April 30). A SARS-CoV-2 protein interaction map reveals targets for drug repurposing [Digital image]. Retrieved January 22, 2021, from https://www.nature.com/articles/s41586-020-2286-9#auth-Zun_Zar_Chi-Naing
Guo, Y., Tian, X., Wang, X., & Xiao, Z. (2018). Adverse effects of immunoglobulin therapy. Frontiers in Immunology, 9(JUN), 1299. https://doi.org/10.3389/fimmu.2018.01299
Gupta, R. (2020). Combating Drug Resistance [PowerPoint]. McMaster University, Hamilton ON.
Hajdo, L., Szulc, A. B., Klajnert, B., & Bryszewska, M. (2010). Metabolic limitations of the use of nucleoside analogs in cancer therapy may be overcome by application of nanoparticles as drug carriers: A review. Drug Development Research, 71(7), 383-394.
Hamming, I., Timens, W., Bulthuis, M., Lely, A., Navis, G., & Goor, H. V. (2004). Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. The Journal of Pathology, 203(2), 631-637. doi:10.1002/path.1570
Hermann, T. (2016). Small molecules targeting viral RNA. Wiley Interdisciplinary Reviews: RNA, 7(6), 726-743.
Huang, Y., Yang, C., Xu, X., Xu, W., & Liu, S. (2020). Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacologica Sinica, 41(9), 1141-1149. doi:10.1038/s41401-020-0485-4
Immunotherapy - StatPearls - NCBI Bookshelf. (n.d.). Retrieved January 29, 2021, from https://www.ncbi.nlm.nih.gov/books/NBK519046/
Jiang, S., Zhang, X., Yang, Y., Hotez, P. J., & Du, L. (2020). Neutralizing antibodies for the treatment of COVID-19. Nature Biomedical Engineering, 4(12), 1134-1139. Retrieved from https://www.nature.com/articles/s41551-020-00660-2
Jin, Z., Du, X., Xu, Y., Deng, Y., Liu, M., Zhao, Y., … & Yang, H. (2020). Structure of M pro from SARS-CoV-2 and discovery of its inhibitors. Nature, 582(7811), 289-293.
Kavianpour, M., Saleh, M., & Verdi, J. (2020). The role of mesenchymal stromal cells in immune modulation of COVID-19: Focus on cytokine storm. Stem Cell Research and Therapy, 11(1). https://doi.org/10.1186/s13287-020-01849-7
Kimiz-Gebologlu, I., Gluce-Iz, S., & Biray-Avci, C. (2018). Monoclonal antibodies in cancer immunotherapy. Molecular Biology Reports, 45, 2935-2940. https://doi.org/10.1007/s11033-018-4427-x
King, J., Kosinski-Collins, M., & Sundberg, E. (2020, March/April). Structure and Organization of Coronaviruses [Digital image]. Retrieved January 22, 2021, from https://web.mit.edu/fnl/volume/324/king_etal.html
King, A., Vail, A., O’Leary, C., Hannan, C., Brough, D., Patel, H., Galea, J., Ogungbenro, K., Wright, M., Pathmanaban, O., Hulme, S., & Allan, S. (2020). Anakinra in COVID-19: important considerations for clinical trials. The Lancet Rheumatology, 2(7), e379–e381. https://doi.org/10.1016/S2665-9913(20)30160-0
Kumar, S., Sharma, V., & Priya, K. (2020). Battle against COVID-19: Efficacy of Convalescent Plasma as an emergency therapy. The American Journal of Emergency Medicine, 0(0). https://doi.org/10.1016/j.ajem.2020.05.101
Liu, J. K.H. (2014). The history of monoclonal antibody development – Progress, remaining challenges and future innovations. Annals of Medicine and Surgery, 3(4), 113-116. https://doi.org/10.1016/j.amsu.2014.09.001
Liu, L., Wang, P., Nair, M. S., Yu, J., Rapp, M., Wang, Q., … & Ho, D. D. (2020). Potent neutralizing antibodies against multiple epitopes on SARS-CoV-2 spike. Nature, 584(7821), 450-456.
Mahmuda, A., Bande, F., Al-Zihiry, K., Abdulhaleem, N., Majid, R., … & Unyah, Z. (2017). Monoclonal antibodies: A review of therapeutic applications and future prospects. Tropical Journal of Pharmaceutical Research, 16(3), 713-722. doi: 10.4314/tjpr.v16i3.29
Manis, J. P. (2020). Overview of therapeutic monoclonal antibodies. UpToDate. Retrieved from https://www.uptodate.com/contents/overview-of-therapeutic-monoclonal-antibodies#H1
Mansourabadi, A. H., Sadeghalvad, M., Mohammadi-Motlagh, H. R., & Rezaei, N. (2020). The immune system as a target for therapy of SARS-CoV-2: A systematic review of the current immunotherapies for COVID-19. In Life Sciences (Vol. 258, p. 118185). Elsevier Inc. https://doi.org/10.1016/j.lfs.2020.118185
Nagarajan, T., Marissen, W. E., & Rupprecht, C. (2014). Monoclonal antibodies for the prevention of rabies: Theory and clinical practice. Antibody Technology Journal, 4(4), 1-12. doi: 10.2147/ANTI.S33533
Nagoba, B., Gavkare, A., Jamadar, N., Mumbre, S., & Selkar, S. (2020). Positive aspects, negative aspects and limitations of plasma therapy with special reference to COVID-19. Journal of Infection and Public Health, 13(12), 1818–1822. https://doi.org/10.1016/j.jiph.2020.08.011
NCBI. (2021, January 3). ORF1ab ORF1a polyprotein;ORF1ab polyprotein [Severe acute respiratory syndrome coronavirus 2]. Retrieved January 22, 2021, from https://www.ncbi.nlm.nih.gov/gene/43740578
Ngo, H. X., & Garneau-Tsodikova, S. (2018). What are the drugs of the future?. MedChemComm, 9(5), 757–758. https://doi.org/10.1039/c8md90019a
Pashaei, M., & Rezaei, N. (2020). Immunotherapy for SARS-CoV-2: potential opportunities. In Expert Opinion on Biological Therapy (Vol. 20, Issue 10, pp. 1111–1116). Taylor and Francis Ltd. https://doi.org/10.1080/14712598.2020.1807933
Pinto, D., Park, Y. J., Beltramello, M., Walls, A. C., Tortorici, M. A., Bianchi, S., … & Corti, D. (2020). Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature, 583(7815), 290-295.
Prescott, H. C., & Rice, T. W. (2020). Corticosteroids in COVID-19 ARDS: evidence and hope during the pandemic. Jama, 324(13), 1292-1295.
Public Health Ontario. (2020, December 16). COVID-19 Vaccines: mRNA Vaccines. Retrieved January 21, 2021, from https://www.publichealthontario.ca/-/media/documents/ncov/vaccines/2020/12/covid-19-mrna-vaccines.pdf?la=en
Ragab, D., Salah Eldin, H., Taeimah, M., Khattab, R., & Salem, R. (2020). The COVID-19 cytokine storm; what we know so far. Frontiers in immunology, 11, 1446.
Ringe, R., & Bhattacharya, J. (2013). Preventive and therapeutic applications of neutralizing antibodies to Human Immunodeficiency Virus Type 1 (HIV-1). Therapeutic advances in vaccines, 1(2), 67-80.
Salazar, G., Zhang, N., Fu, T. M., & An, Z. (2017). Antibody therapies for the prevention and treatment of viral infections. npj Vaccines, 2, 19. https://doi.org/10.1038/s41541-017-0019-3
Shereen, M. A., Khan, S., Kazmi, A., Bashir, N., & Siddique, R. (2020). COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. Journal of Advanced Research, 24, 91-98. doi:10.1016/j.jare.2020.03.005
Smieszek, S., Przychodzen, B., Polymeropoulos, V., Polymeropoulos, C., & Polymeropoulos, M. (2020). Direct ACE2-Spike RBD Binding Disruption with Small Molecules: A Strategy for COVID-19 Treatment.
Tang, X., Wu, C., Li, X., Song, Y., Yao, X., Wu, X., . . . Lu, J. (2020). On the origin and continuing evolution of SARS-CoV-2. National Science Review, 7(6), 1012-1023. doi:10.1093/nsr/nwaa036
Tortorici, M. A., Beltramello, M., Lempp, F. A., Pinto, D., Dang, H. V., Rosen, L. E., … & Veesler, D. (2020). Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms. Science, 370(6519), 950-957.
Walls, A. C., Park, Y. J., Tortorici, M. A., Wall, A., McGuire, A. T., & Veesler, D. (2020). Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell, 181(2), 281-292.
WHO. (2020, June 25). Coronavirus disease (COVID-19): Dexamethasone. Retrieved January 22, 2021, from https://www.who.int/news-room/q-a-detail/coronavirus-disease-covid-19-dexamethasone
Wraith, D. C. (2017). The future of immunotherapy: A 20-year perspective. Frontiers in Immunology, 8(NOV), 1668. https://doi.org/10.3389/fimmu.2017.01668
Wu, Y., Li, Z., Zhao, Y., Huang, Y., Jiang, M., & Luo, H. (2021). Therapeutic targets and potential agents for the treatment of COVID-19. Medicinal Research Reviews, 1-23. doi:10.1002/med.21776
Yi, L., Li, Z., Yuan, K., Qu, X., Chen, J., Wang, G., … & Xu, X. (2004). Small molecules blocking the entry of severe acute respiratory syndrome coronavirus into host cells. Journal of virology, 78(20), 11334-11339.
Zost, S. J., Gilchuk, P., Chen, R. E., Case, J. B., Reidy, J. X., … & Crowe Jr, J. E. (2020). Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein. Nature Medicine, 26, 1422-1427. https://doi.org/10.1038/s41591-020-0998-x